We are proud to release a new version of VarSome introducing two new features which we hope you will find useful.
ACMG Classification
An automated classification tool based on the ACMG guidelines [1]. This is an experimental tool for educational purposes only. It classifies the variant automatically using all the data available to VarSome, following our interpretation of the ACMG guidelines. Clicking on the blue question marks will give information about the rule itself and any matching criteria. The section below it proves an English description of the evidence. Users can toggle findings on or off, if they disagree or if they have additional data. The overall verdict will update accordingly.
Lollipop Graph
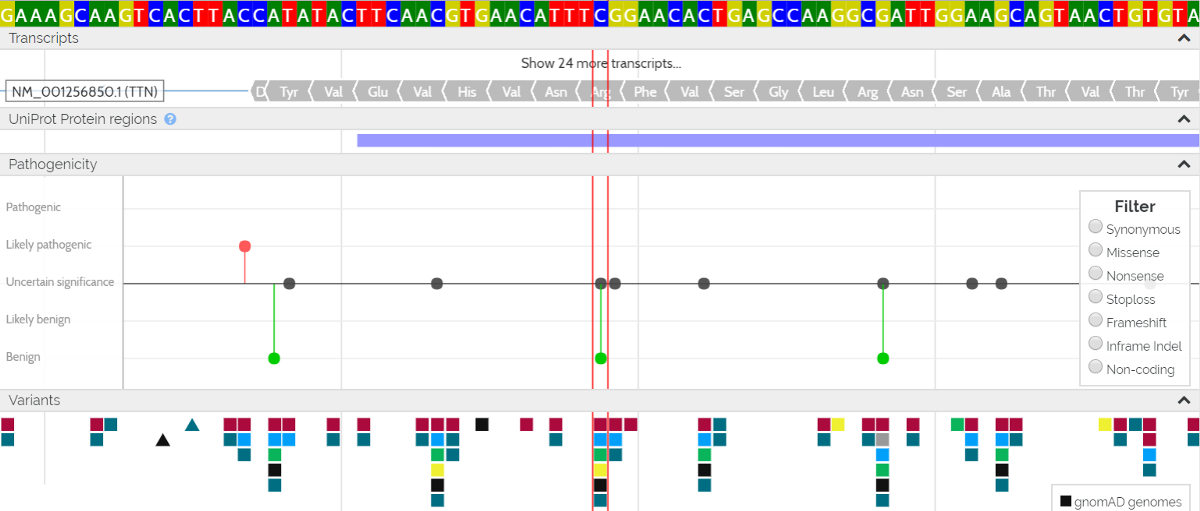
A “Lollipop” graph shows all the known variants in ClinVar, UniProt or contributed by VarSome users. You can hover over any of the dots to see more information on the variant, or click on a variant to select it. The main aim of this graph is to provide a high-level genomic view of where the pathogenic and benign variants are to be found.
References
- [1] Richards S. et al, Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology, Genetics in Medicine 2015.

